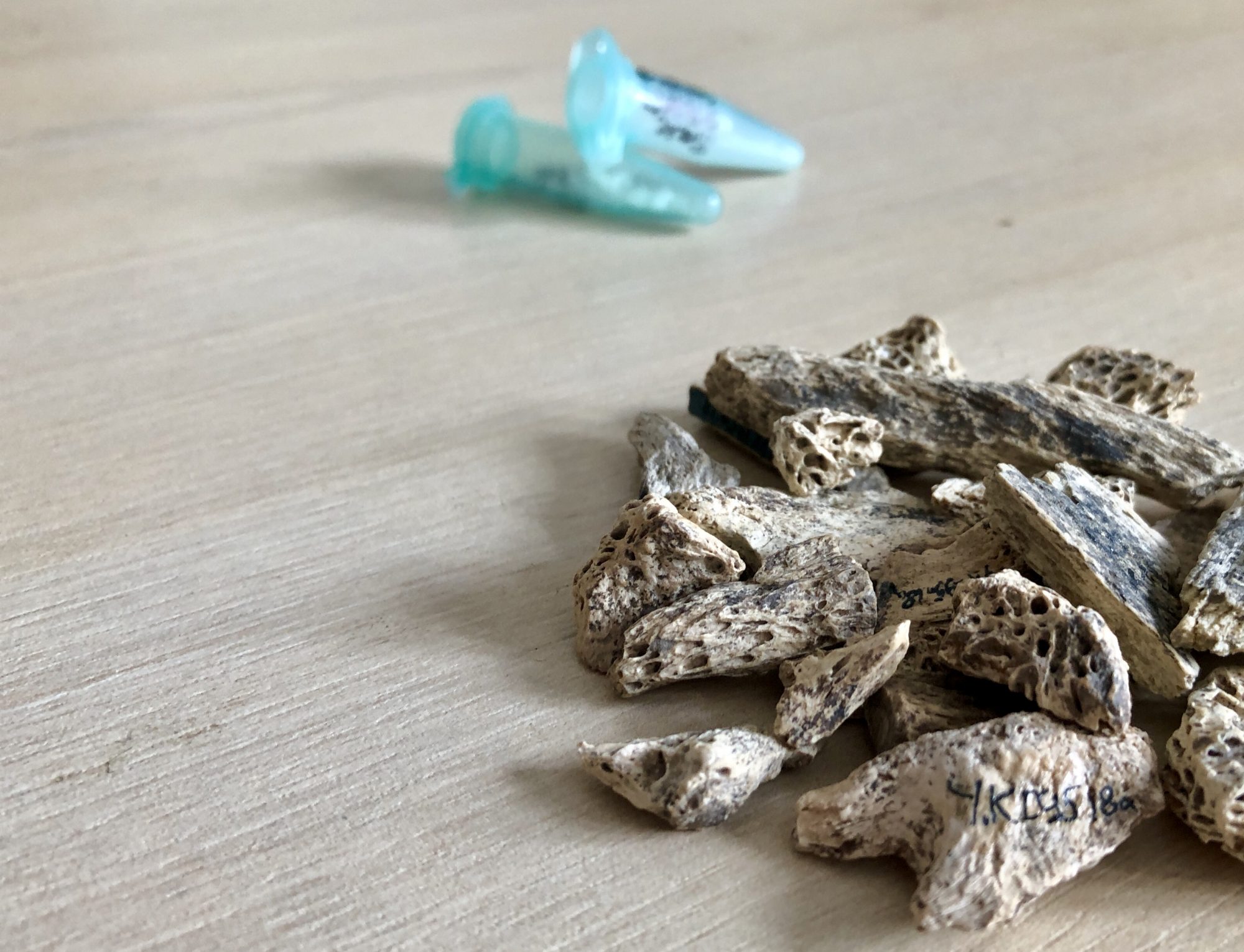
This biomolecular method, developed for the analysis of bone collagen about a decade ago by M. Collins and M. Buckley at the University of York, is a fast and cheap screening approach for a first-level taxonomic identification of bones.
Collagen is the dominant, and often, best preserved protein in tissues such as bone, dentine and antler. From a few milligrams of powder collagen type I (COL1) is extracted and digested to peptides, and the peptides are analysed using mass spectrometry (MALDI-ToF-MS) to produce a collagen peptide mass fingerprint. This fingerprint is unique for deeply-diverged taxonomic groups, hence the observed spectrometric patterns are compared and matched against a reference database consisting of known-origin peptide fingerprints.

In this project we use collagen peptide mass fingerprinting as a first step identification of bones belonging to the Hominidae. The fragments are then CT-scanned and photographed before proceeding to the second level of specialised analyses which consist of:

